Afr. J. Parasitol. Mycol. Entomol. , 3(2), 13; doi:10.35995/ajpme03020013
Article
Assessment of Malaria Infection Prevalence, Parasite Genetic Diversity, and Prevalence of Plasmodium falciparum Resistance Markers During Seasonal Malaria Chemoprevention in Children Living in Saponé, Burkina Faso
1
Biomedical and Public Health Department, Institut de Recherche en Sciences de la Santé (IRSS), Centre National de 03, BP 7047 Ouagadougou 03, Burkina Faso; soulamacnrfp@gmail.com (I.S.); ouedoumarou.77@gmail.com (O.O.); zabdouazaque@cnrst.gov.bf (A.A.Z.); ilyraissa55@gmail.com (R.I.); tibysnagui@gmail.com (G.Y.N.T.); zoureaicha6@gmail.com (D.O.A.Z.); yanogonas@gmail.com (J.N.Y.); zdramane@gmail.com (D.Z.)
2
Research Department, Centre National de Recherche et de Formation sur le Paludisme (CNRFP), Institut National de Santé Publique (INSP), BP 10278 Ouagadougou, Burkina Faso; salifsombie@yahoo.fr (S.S.); nouedraogo94@gmail.com (N.O.); faridakabore29@gmail.com (F.C.A.K.)
3
Laboratoire de Biologie Moléculaire et de Génétique (LABIOGENE), Université Joseph KI-ZERBO, Ouagadougou, Burkina Faso; florencia.djigma@gmail.com (F.W.D.); simpore93@gmail.com (J.S.)
4
Laboratoire d’Entomologie Fondamentale et Appliquée (LEFA), Université Joseph KI-ZERBO, Ouagadougou, Burkina Faso; moudiammoungou@gmail.com (A.A.M.); a.badolo@gmail.com (A.B.)
5
Direction Scientifique, Groupe de Recherche Action en Santé, Ouagadougou, Burkina Faso; sermesame@yahoo.fr
6
Unité de Formation et de Recherche en Sciences de la Santé (UFR/SDS), Université Joseph Ki-ZERBO, Ouagadougou, Burkina Faso; ephiaouare@gmail.com (N.O.); salamsawadog19@gmail.com (S.S.); zidaadama@live.fr (A.Z.)
7
Unité de Recherche Clinique de Nanoro (URCN), Nanoro, Burkina Faso
8
Service de Parasitologie-Mycologie, Centre Hospitalier Universitaire Yalgado Ouedraogo (CHU-YO), Ouagadougou 03, Burkina Faso; haffsasawa@yahoo.fr
9
Laboratoire de Biochimie et Immunologie Appliquées (LABIA), Université Joseph KI-ZERBO, Ouagadougou, Burkina Faso; chanoltchek@gmail.com
10
Institut International des Sciences et Technologie (IISTech), Ouagadougou, Burkina Faso
11
Laboratoire de Biologie et d Ecologie Animale (LBEA), Université Joseph KI-ZERBO, Ouagadougou, Burkina Faso
12
Centre de Recherche Biomoléculaire Pietro Annigoni (CERBA), Ouagadougou, Burkina Faso
*
Corresponding author: seninikiema@yahoo.fr or sjseninikiema@gmail.com; Tel.: +226-63709155 or +226-74784434
How to cite: Soulama, I.; Nikiema, S.; Ouedraogo, O.; Sombié, S.; Ouedraogo, N.; Zouré, A.A.; Sermé, S.S.; Ouaré, N.; Sawadogo, S.; Sawadogo, H.; et al. Assessment of Malaria Infection Prevalence, Parasite Genetic Diversity, and Prevalence of Plasmodium falciparum Resistance Markers During Seasonal Malaria Chemoprevention in Children Living in Saponé, Burkina Faso. Afr. J. Parasitol. Mycol. Entomol. 2026, 3(2): 13; doi:10.35995/ajpme03020013.
Received: 15 June 2025 / Accepted: 12 March 2026 / Published: 8 May 2026
Abstract
:Background: Seasonal Malaria Chemoprevention (SMC), consisting of a combination of sulfadoxine pyrimethamine (SP) and amodiaquine (AQ), aims to reduce morbidity and mortality, in children under five years of age. The present study aims to assess Plasmodium falciparum’s level of genetic diversity and the dynamics of parasite-resistance-associated mutations during SMC implementation in children living in Saponé, Burkina Faso. Methods: The study included 222 children aged 3 to 59 months under SMC in the Saponé health district. Parasite DNA was extracted from the blood spots using the QIAamp® DNA Blood Mini Kit, and genetic diversity was determined by nested PCR (Polymerase Chain Reaction) of P. falciparum msp-1 and msp-2 genes, while molecular markers of resistance were analyzed based on PCR/RFLPs. Results: The molecular results showed that the prevalence of malaria infection by PCR averaged 17.43% ± 0.30 during the SMC. The study revealed a high genetic diversity of the parasite during SMC. The value of the complexity of the msp-1 and msp-2 genes decreased during the SMC rounds, reflecting a reduction in malaria transmission during this strategy. The prevalence of single-nucleotide polymorphism mutations associated with SP+AQ (sulfadoxine–pyrimethamine + amodiaquine) resistance was very low or nil during our study. However, the prevalence of the Pfdhfr59 mutation was very high, standing at an average of 91. Conclusions: The prevalence of mutations in genes associated with parasite resistance to SP+AQ was low in the study population during SMC. Therefore, the sulfadoxine–pyrimethamine drug combination used in SMC still remains appropriate for the treatment of malaria in children under five years old in Burkina Faso.
Keywords:
Malaria; Plasmodium falciparum; polymorphism; SMC; resistance markers; Saponé; Burkina Faso1. Background
Malaria is the deadliest parasitic infection and is transmitted to humans through the bites of infected female Anopheles mosquitoes during their blood meal. Approximately 200 species of Plasmodium infect mammals, rodents, birds and amphibians, and at least five species of Plasmodium are responsible for malaria in humans. Of these species, Plasmodium falciparum is the most widespread, the most dangerous, and is responsible for often fatal complications [1]. Despite considerable progress in control, this disease remains a public health problem in many regions, particularly in sub-Saharan Africa. In 2024, estimates suggested that there would be 11 million more cases of malaria in 2023 than in 2022 (263 million versus 252 million), with most of this increase occurring in the African region countries [2]. Indeed, in sub-Saharan Africa, the estimated number of malaria deaths increased by 12% in 2020 compared to 2019. Also, there has been a worldwide increase in the number of deaths related to malaria, with 69,000 more people dying from malaria in 2020 compared to 2019 (627,000 against 558,000). Almost two-thirds (47,000) of these additional malaria deaths resulted from disruptions in malaria prevention, diagnosis, and treatment services during the COVID-19 pandemic [3]. These figures highlight the consequences of even moderate disruptions to malaria services in a population at risk. More than 80% of deaths are among children under five and during periods of high malaria transmission. Seasonal Malaria Chemoprevention (SMC) was recommended in 2012 by the WHO to reduce morbidity and mortality during periods of high transmission in endemic areas in Africa. This strategy uses sulfadoxine–pyrimethamine (SP) in combination with amodiaquine (AQ) and is an effective and promising strategy for controlling malaria in areas characterized by high seasonal malaria transmission [4]. Implementation of SMC in Burkina Faso began in 2014, along with several other African countries. First implemented in endemic areas, it is now being implemented countrywide. The results of a recent study showed that SMC is a high-potential intervention in Burkina Faso. In conjunction with other interventions, it could contribute to achieving elimination strategies for this disease [5]. However, based on national statistical data [6], the number of malaria cases in the target group increased by 22% between 2016 and 2017.
The increase in the prevalence of infection and the pressure to use SP+AQ since the implementation of SMC therefore suggests the importance of integrating molecular surveillance, particularly genetic diversity and parasite resistance markers. This diversity and these markers of parasite resistance strongly correlate with the dynamics of the level of transmission and could be associated with therapeutic failures.
The current study was initiated to evaluate the level of genetic diversity and the prevalence of P. falciparum resistance markers during SMC in children living in Saponé, Burkina Faso.
2. Materials and Methods
2.1. Study Area
The study was conducted in Saponé’s health district (Figure 1), located in the province of Bazéga, a rural area 50 km southwest of Ouagadougou, the capital of Burkina Faso, between 12°13′ N and 1°48′ W, where malaria is mesoendemic. Malaria transmission in this district is permanent, with a peak during the intense, seasonal rainy season from June to November. Chemoprevention of seasonal malaria has been implemented since 2014 in this area.
Figure 1.
Study site, Burkina Faso, 2020.
2.2. Study Population and Sample Collection
This study involved 222 male and female children aged 3 to 59 months receiving SP+AQ as part of Seasonal Malaria Chemoprevention. Participants were recruited, following a random selection methodology, from villages located in the Saponé health district area. These participants were reviewed during the malaria transmission season and concurrently with the four rounds of the 2020 SMC campaign from July to October. For each patient included in the study, blood samples were taken via finger prick to prepare blood spots on filter paper and blood smears for malaria diagnosis. The filter paper spots were kept in a plastic bag containing silica gel to keep the medium dry, before being inserted into an envelope and transported to the molecular biology laboratory at CNRFP (Centre National de Recherche et de Formation sur le Paludisme) for molecular analysis.
2.3. Asexual Plasmodium falciparum Parasite Density and Hemoglobin Level Determination
Thick and thin blood smears were used for microscopic diagnosis of parasite count in the parasitology laboratory at CNRFP. Thick and thin blood smears were stained with Giemsa. The number of asexual P. falciparum parasites was estimated for 200 leucocytes, and parasite density was calculated assuming a mean of 8000 leucocytes per μL of whole blood. A sample was declared negative after examining 200 thick-film fields without observing any asexual parasite.
2.4. Plasmodium falciparum DNA Extraction
Plasmodium falciparum DNA was extracted from the blood spots using Qiagen® DNA extraction kits according to the manufacturer’s instructions (https://www.iddo.org/procedure/dna-extraction-qiaamp-dna-mini-kit, accessed 20 September 2025) and stored at −20 °C until use.
2.5. Plasmodium falciparum Identification by Nested PCR
The nested PCR method was used for the amplification of Plasmodium falciparum DNA sequences. This PCR variant consists of a consecutive series of two PCRs using two different primer pairs (1st amplification and 2nd amplification). The first amplification was performed in a 20 μL reaction volume consisting of 4 μL of 5X FIREPol® Master Mix Ready to Load, 1 μL of Plu5 primer (5′ CCTGTTGTTGCCTTAAACTTC 3′) at 10 μM, 1 μL Plu6 (5′ TTAAAATTGTTGCAGTTAAAACG 3′) at 10 μM, 13.5 μL of distilled water, and 0.5 μL of parasite DNA to amplify a large portion of the variable region of the desired P. falciparum gene. The second amplification was performed using the product of the first, with the same reaction volume and the same components, except for the primer pair Fal1 (5′ TTAAACTGGTTTGGGAAAACCAAATATATT3′) and (5′ACACAATGAACTCAATCATGACTACCCGTC3′) Fal2 at 1 μL for each primer to amplify a more specific region of the gene. This primer pair (Fal1 and Fal2 at 1 μL) is specific to the region containing the P. falciparum gene. After the second PCR amplification, electrophoresis on a 2% agarose gel stained with Redsafe, a fluorescent intercalant, was performed. The amplified DNA containing various clones was then visualized by a UV device. The sample containing a band of 205bp is considered positive for Plasmodium falciparum.
2.6. Molecular Genotyping
2.6.1. Search for msp-1 and msp-2 Polymorphisms
The search for polymorphisms on block 2 of msp-1 and block 3 of msp-2 was performed via nested PCR using two consecutive PCRs and using specific primer pairs according to a previously described protocol, with slight modifications [7]. The first nested PCR used a primer pair (sense, antisense) to amplify a large part of the variable region of the desired gene. The second was performed using the product of the first—with the same reaction volume and the same components, except for the primer pair (sense, antisense)—to amplify a more specific region of the desired gene. A sample was considered to belong to a given allelic family (msp-1 and msp-2) if there was the appearance of at least one band visualized by the ultraviolet transilluminator after electrophoresis on the 2% agarose gel. The presence of multiple fragments or clones of P. falciparum was considered a polyclonal infection, and a single allele was considered a monoclonal infection. The complexity of infection (COI) corresponds to the average number of clones or strains of P. falciparum in an individual. It is calculated as follows: sum of P. falciparum strains presents in infected subjects/number of infected subjects.
2.6.2. Genotyping of Markers Associated with AQ+SP Resistance by PCR/RFLPs
Evaluation of parasite resistance markers to AQ+SP was also performed by nested PCR followed by enzymatic restriction digestion. The parasite resistance markers analyzed in this study were Pfdhfr N51I, Pfdhfr C59R, Pfdhfr S108N, and Pfdhfr I164L for pyrimethamine; Pfdhps A437G and Pfdhps K540E for sulfadoxine; and Pfcrt K76T and Pfmdr1N86Y for amodiaquine [8]. Characterization of the mutations was also performed using specific enzymes and primers previously described in the University of Maryland protocols by Dr. C. Plowe (http://medschool.umaryland.edu/malaria/protocols/, accessed 20 September 2025) [8], with slight modifications. Briefly, two consecutive amplifications were performed in a 25 μL reaction volume consisting of 4 μL of 5X FIREPol® Master Mix containing (Buffer, dNTP, Mgcl2, Taq polymerase), 1 μL of sense primer (10 μM), 1 μL of antisense primer (10 μM), 13.5 μL of distilled water, and 0.5 μL of parasite DNA. These different amplifications (first and second) followed a unique program (initial denaturation at 95 °C for 5 min:1 cycle; denaturation at 95 °C for 30 s; hybridization at 60 °C for 45 s; elongation at 72 °C for 2 min: 30 cycles; and final elongation at 72 °C for 10 min, before storage at 4 °C), and these reactions were carried out in adherence to the protocol described at https://solisbiodyne.co/pics/6203 (accessed 20 September 2025) where one can find the Data Sheet 5x FIREPol Master_Mix 12.5 Mm MgCl2.pdf of the Applied Biosystem 2720 thermal cycler (Part n°4369669 Serial N° 27250192363). The products of the second PCR were used for enzymatic digestion for resistance mutation detection. This digestion was therefore performed using enzymes specific to each mutation to be analyzed based on the principle of Restriction Fragment Length Polymorphisms (RFLPs). Positive and negative controls were included in each reaction set. Ten microliters of each PCR product were separated by electrophoresis on a 1.5% agarose gel stained with ethidium bromide. The different fragments of the genes (wild-type and mutant) on Pfdhfr and Pfdhps correspond to the fragment sizes given.
2.7. Ethical Approval
The research protocol for this study received approval from the Health Research Ethics Committee of Burkina Faso (Deliberation No. 2020-5-092). The decision to participate in the study was the sole responsibility of the child’s parent or guardian. All study documents (including our data codes) were kept in a secure location; blood samples were labeled with a unique code without the patient’s name to ensure confidentiality.
2.8. Statistical Analysis
The data were entered and analyzed using EXCEL 2020 and STATA version 13, respectively. EXCEL 2020 was also used for graphical representations. The chi-square test was used for comparison of proportions and means. The threshold for statistically significant differences were set at p ˂ 0.05.
3. Results
3.1. Characteristics of Study Population
The study involved 222 children aged 3 to 59 months at the time of first visit in six villages of the Saponé health district, namely Banabenbato, Watenga, Damzoussi, Kontenga, Sambsin, and Tanghin. The study population was 13.96% (31/222) female and 86.04% (191/222) male, giving a sex ratio (M/F) of 6.16 in favor of males. The geometric mean of parasite density was 2581.72 parasites/µL; 5792.97 parasites/µL; 4937.02 parasites/µL; and 4112.40 parasites/µL from the first to the fourth round, respectively. This average remained stable after the different treatments during the SMC. Axillary temperature was evaluated at 36 °C. No significant difference was observed when comparing these clinical and parasitological parameters according to the different passages (p > 0.05). Table 1 (below) presents average age and average temperature data, stratified by gender.
Table 1.
Characteristics of study participants.
| Male | Female | p | |
|---|---|---|---|
| 86% (191/222) | 14% (31/222) | <0.05 | |
| Average age (Months) | 29.75 | 35.71 | <0.05 |
| Average temperature (°C) | 36.40 | 36.04 | >0.05 |
3.2. Prevalence of P. falciparum Infection
Blood samples that were tested and confirmed to be positive for P. falciparum by nested PCR were used for the characterization of parasite polymorphisms and mutation detection from associated genes with SP-AQ resistance. Plasmodium falciparum infection was estimated at 6%, 3%, 30%, and 38% from the first to last round at day 0, giving an overall mean PCR malaria prevalence of 19.25% in our study.
3.3. Frequency of msp-1 and msp-2 Allelic Families
The frequencies of the allelic families of the msp1 and msp2 genes were determined during the different SMC runs in our study (Figure 2). The frequencies of the k1 and ro33 alleles of msp1 were predominant in the first and last two rounds, respectively (69.23%; 83.33% and 98.15%; 98.18%). The frequency of the ro33 allele increased significantly over the four SMC rounds. However, the frequencies of the k1 and mad20 allelic families were not statistically different (p > 0.05) over the rounds despite a decreasing trend (Figure 2a). As for the msp2 allelic families, the frequencies of 3D7 and FC27 were comparable, standing at 72.73% and 67.64% (p = 0.8), respectively, for the first run of the SMC. These FC27 and 3D7 frequencies were predominant from the second run onwards. No statistically significant difference was observed across passages (p = 0.6); however, the 3D7 frequency generally increased, while FC27 decreased during SMC. For data on the msp2 gene allelic families, see Figure 2b.
Figure 2.
Allele frequencies of the msp1 (a) and msp2 (b) genes in the population during SMC.
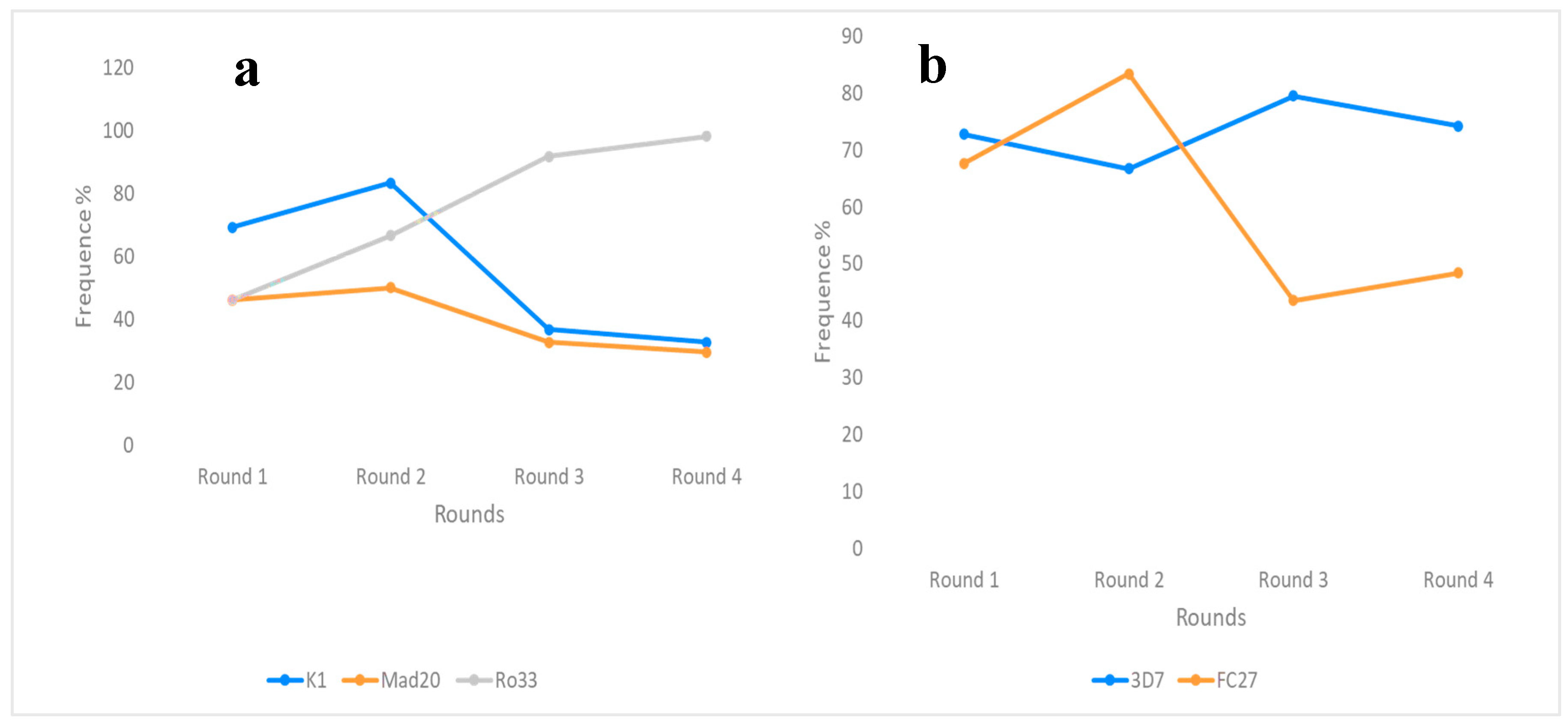
Individual alleles for the msp-1 and msp-2 genes were analyzed in the parasite population according to base pair size. For the msp-1 gene, individual k1 alleles were the most diverse, with eight (8) individual alleles identified compared to mad20 and ro33, which had five (5) and four (4) strains, respectively; thus, we found a total of seventeen different strains for the msp1 gene. Twenty-three (23) different fragments of msp-2 were characterized, including ten different fragments for the FC27 allele and thirteen (13) fragments for the 3D7 allelic family.
3.4. Complexity and Polyclonality of Infection During SMC
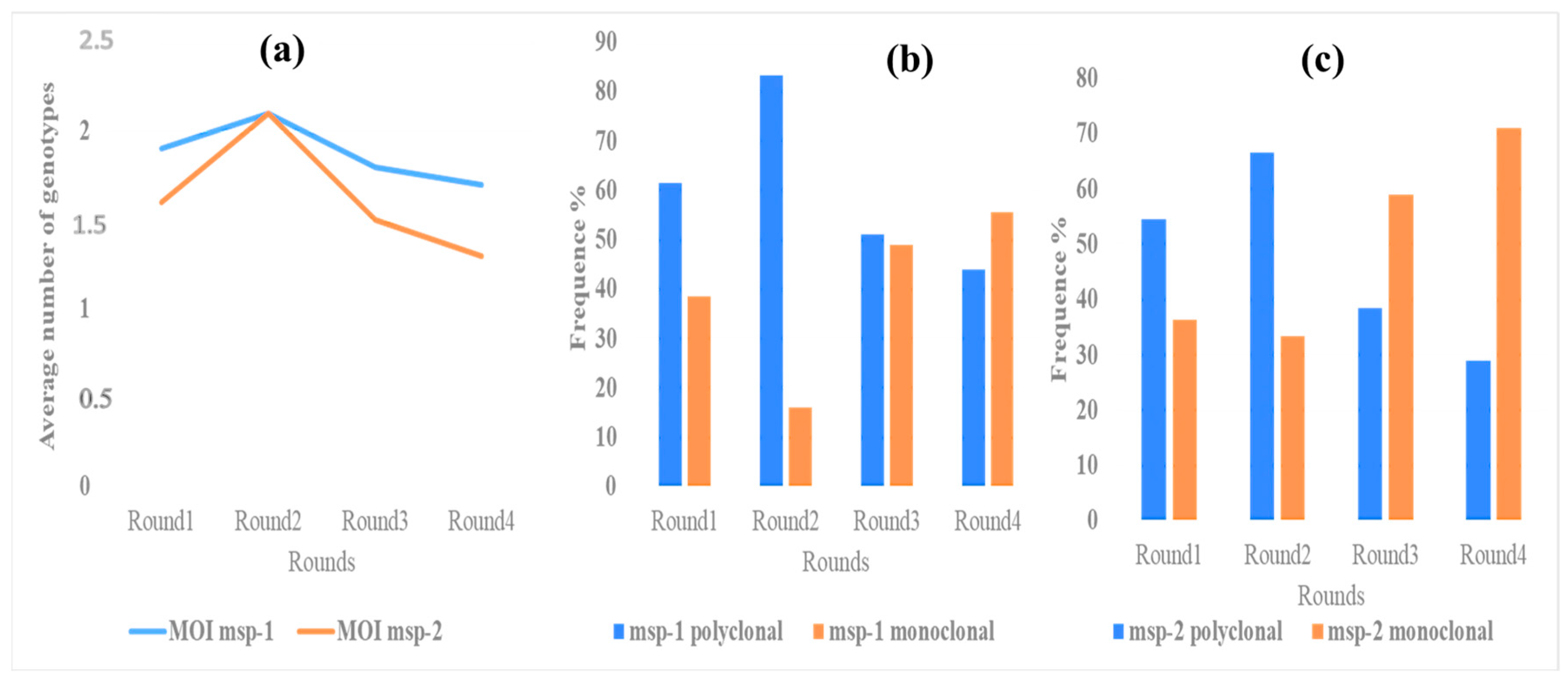
The frequency of polyclonality, defined as the proportion of children with more than one clone (allele) of Plasmodium falciparum at msp1, was 61.54%, 83.33%, 51.02%, and 44% from the first to the last passage of the SMC, respectively. As for the msp-2 gene, polyclonal-type infections were estimated at 63.64%, 66.67%, 38.46%, d 29% from the first to the last round, respectively. This polyclonality of msp-1 and msp-2 was not statistically different during SMC (p > 0.05) and tended to decrease. In our study, the complexity of infections for msp-1 and msp-2 were (1.9; 2.1; 1.8; 1.7) and (1.6; 2.1; 1.5; 1.3) from the first to the last round, respectively. We observed a peak in the complexity of msp-1 and msp-2 gene infections in the second round. However, a downward trend in the complexity of msp-1 and msp-2 gene infections was observed from the third round to the fourth round. There was no significant difference in the evolution of the complexity of infection (COI) between the first and fourth rounds (p > 0.05). These data are shown in Figure 3.
Figure 3.
Evolution of complexity (a) and polyclonality of msp-1 (b) and msp-2 (c) during SMC.
3.5. Prevalence of Resistance Genes Associated with SP+AQ Resistance
The frequencies of mutations in the genes associated with parasite resistance to QA and SP (Pfcrt and Pfmdr1 and Pfdhfr and Pfdhps, respectively) were assessed during SMC. The prevalence of the PfcrtK76T and Pfmdr1-N86Y mutations was very low or nil during SMC in our study. Indeed, the PfcrtK76T frequency was null during the first two passages. However, the prevalence of PfcrtK76T mutations was 6.12% and 7.27, respectively, for passages 3 and 4. Pfdhfr 51,108,164; Pfdhps 437,540; Pfdhfr (51, 59); Pfdhfr (51, 108); Pfdhfr (59, 108); and Pfdhfr (51, 59, 108) triple mutations—as well as Pfdhfr (51, 59, 108) and Pfdhps (437, 540) quintuple mutations—had very low or negligible or null proportions (Table 2). However, the prevalence of mutation on codon 59 of the Pfdhfr gene was high in all four passages (>83.33%).
Table 2.
Prevalence of SNPs in genes associated with SP+AQ resistance.
| Gene Mutation | Round 1 | Round 2 | Round 3 | Round 4 | p |
|---|---|---|---|---|---|
| PfcrtK76T | 0 (0%) | 0 (0%) | (6.12%) | (7.41%) | >0.05 |
| Pmdr1-N86Y | 1 (7.69) | 0 | 0 | 0 | |
| Pfdhfr N51I | 1 (7.69%) | 1 (16.67%) | 1(2.04%) | 1 (1.85%) | |
| Pfdhfr C59R | 12 (92.31%) | 5 (83.33%) | 46 (93.88%) | 51 (94.44%) | |
| Pfdhfr S108N | 0 (0%) | 0 (0%) | 6 (12.24%) | 0(0%) | |
| Pfdhfr I164L | 0 (0%) | 0 (0%) | 2 (4.08%) | 1 (1.84%) | |
| Pfdhps A437G | 2 (15.36%) | 0 (0%) | 1 (2.04%) | 3 (5.56%) | |
| Pfdhps S540E | 0 (0%) | 0 (0%) | 0(00%) | 1 (1.85%) | |
| Pfdhfr (51, 59) | 1 (7.69%) | 1 (16.67%) | 1 (2.04%) | 1 (1.85%) | |
| Pfdhfr (51, 108) | 0 (0%) | 0 (0%) | 1 (2.04%) | 0 (0%) | |
| Pfdhfr (59, 108) | 0 (0%) | 0 (0%) | 1 (2.04%) | 0 (0%) | |
| Pfdhfr (51, 59, 108) | 0 (0%) | 0 (0%) | 1 (2.04%) | 0 (0%) | |
| Pfdhfr (51, 59, 108) dhps437-540 | 0 (0%) | 0 (0%) | 0 (0%) | 0 (0%) |
4. Discussion
Plasmodium falciparum has a high genetic polymorphism that confers the ability to develop multidrug resistance or to bypass vaccines. The emergence and spread of resistant strains is an obstacle in the fight against malaria [9]. A better knowledge of the circulating strains of P. falciparum would be an important element in the adjustment of control strategies for this parasite [10]. Furthermore, this genetic diversity determines the level of malaria transmission, providing baseline data for any drug efficacy trials and the ability to implement vaccine-based control methods. The surface proteins msp-1 and msp-2 are widely used to study allelic diversity and frequency of P. falciparum highly correlated with the level of transmission [11]. They also represent a discriminatory tool for the distinction between new and recrudescent infections [12] during drug efficacy studies. The msp1 and msp2 genes have three allelic families (mad20, k1, and ro33) and two allelic families (3D7 and FC27), respectively. Our study showed an increase and decrease in the prevalence of the ro33 and (k1, mad20) alleles, respectively. The k1 and ro33 families were predominant in the first two rounds and the last two rounds, respectively. However, studies have shown that the k1 allelic family can be predominant, followed by the mad20 allelic family, with values of 77.4% and 100%, respectively [13]. This variability indicates that the two genes, msp-1 and msp-2, are highly polymorphic in P. falciparum strains, especially those in Africa, with some variations related to malaria epidemiological areas [14]. Various studies have shown that the genetic diversity of P. falciparum populations varies according to the intensity of malaria transmission in endemic areas and that it is only higher in hyperendemic areas than in low-endemicity areas [15]. Thus, a limitation of this study is the fact that we measured the genetic diversity of parasite populations by using markers based on DNA band size. This is a limitation because fragments of the same size (not identical) may be considered identical, giving a false impression of similarity. Within allele families, alleles of the same size may have different amino acid profiles. Indeed, it has been reported that alleles of the same size and allele family are not necessarily identical [16]. Nevertheless, these msp-1 and msp-2 gene markers remain good polymorphism markers, and they can be successfully used to characterize populations of genetic strains of P. falciparum. Allelic variation and the presence of common alleles could be better explored by sequencing.
The two aforementioned markers (msp-1 and msp-2) allow for the establishment of a relationship between parasite polymorphism and the most commonly observed and measurable signs of malaria, such as temperature, parasitemia, and hemoglobin levels [17]. Several studies have shown an association between the prevalence of certain clones, allelic families of the parasite, and clinical signs of disease [7,12,17]. The mad20 and k1 allelic families are thought to increase the risk of developing symptomatic malaria [18]. In Gabon, a significant association between the mad20 allelic family and malaria severity was observed by Bouyou-Akotet et al. in 2015 [19]. A study conducted in France by Guiana by Ariey et al. showed an overrepresentation of msp1 (B-K1) alleles that was much greater in severe malaria than in simple malaria, similar to Ariey et al.’s research from 2001 [20]. Our study showed a decrease in mad20 and k1, and this could therefore represent an advantage for children positive for P. falciparum infection.
As for the msp2 gene, the severity of malaria is related to the prevalence of specific clone types as well. In our study, FC27 and 3D7 frequencies were predominant in the second and last two rounds, respectively. The 3D7 frequency generally increased, while FC27 decreased during the SMC. This decrease in FC27 prevalence could also be advantageous for P. falciparum-positive children. Indeed, one study has shown that the FC27 allelic family is significantly more associated with low hemoglobin values [17]. A study conducted in Papua New Guinea showed this association between the msp-2-FC27 parasite carriage and disease severity [21]. Concerning the msp2-3D7 allelic family, a study in Gabon showed a higher frequency of the 3D7 allelic family in asymptomatic infections compared to symptomatic infections. Predominance of 3D7 genotypes was also observed in severe cases without severe anemia when compared to severe cases with severe anemia [22]. However, analysis of the results revealed that the frequency of 3D7 parasite forms was not different between severe and non-severe cases [14].
According to numerous studies conducted in different localities, knowledge on the association between parasite genetic diversity and malaria symptomatology remains limited due to discrepancies in findings [17]. Polyclonality represents the proportion of children with more than one P. falciparum clone, and this polyclonality determines the degree of complexity, defined as the average number of parasite genotypes present in a subject. The higher the polyclonality in subjects, the higher the complexity of the disease. The results of this study show a reduction in polyclonality during SMC and therefore indicate a decrease in the multiplicity of infections of the msp-1 and msp-2 genes, which would represent an advantage in treatment. Complexity is also related to the severity of P. falciparum infection, because multiple infections (especially those of high complexity) could reflect a failure of immune protection. The presence of several distinct clones at once results in an overload of the immune response that becomes ineffective [23,24]. Results from one study also showed that high complexity can contribute to the severity of malaria anemia [25]. The variability in the complexity of malaria infection, as well as the impact of the environment on the biology of the parasite and the mosquito, makes it quite difficult not only to control malaria but also to generalize the results, hence the need for better knowledge of both local and regional genetic diversity of the parasite and the vector [14].
Resistance to AQ and its metabolite has been linked to Pfcrt76T and Pfmdr1-N86Y mutations [22,26]. In our study, the prevalence of the Pfcrt76T mutation is null during the first round and sets in from the third round onwards. The prevalence of the Pfmdr1-N86Y mutation was 7.67% in the first round and null for the three subsequent rounds. The prevalence of these Pfmdr1-N86Y and Pfcrt76T mutations remained low overall in our study, thus indicating that SMC remains appropriate for use in the treatment and prevention of malaria in children under five during the period of high transmission. This occurrence of Pfcrt76T mutation could be explained by non-compliance with the normal indicated dose, as well as the treatment duration, which could promote mutations associated with drug resistance [27,28]. Most mothers in rural settings are illiterate and have difficulty assimilating instructions on how to take drugs given by health workers [4,29]. It has also been shown that parasites combining both Pfcrt76T and Pfmdr1-N86Y mutations are resistant to 4-aminoquinolines and associated with clinical failure of AQ treatment [27], with a possible negative impact on the efficacy of SMC. In our study, we did not find strains combining both mutated genes (PfcrtK76T+Pfmdr1-N86Y). Also, in a recent study (meta-analysis), the Pfmdr1-N86Y allele was identified as an independent risk factor for occurrence of recrudescence, and therefore, this selective pressure should be carefully monitored [30]. In such a context, even if the selected strains are not associated with treatment failures, this could compromise the efficacy of SP+AQ in the future (when mutant strains will be predominant in the area) by decreasing susceptibility to AQ in areas that have experienced high chloroquine resistance, such as Burkina Faso.
In this study, only a high prevalence of Pfdhfr59 was recorded—92.31%, 83.33%, 93.88%, and 94.44% during round 1, round 2, round 3, and round 4, respectively. A high prevalence of this Pfdhr59 mutation during SMC implementation was found 99.9% of participants in a study conducted in Mali in 2014, and almost all samples (98.8%) in children carried the PfdhfrC59R mutation at baseline [31]. This is why the author states that this mutation is no longer relevant for monitoring sulfadoxine–pyrimethamine resistance. These results are consistent with those obtained at other sites in Mali and other Sahelian countries [32,33,34]. The other Pfdhfr (51, 108, 164) and Pfdhps (437, 540) mutations were very poorly represented, negligible, or absent in our study. Our samples had very few triple mutations (0.25%) and no quintuple mutations. The quintuple mutation is known to be strongly associated with clinical failure of therapy [32]. PfdhpsS540E and PfdhpsA437G mutation prevalence is most commonly used to monitor P. falciparum’s resistance to sulfadoxine. On the other hand, the role of mutations at certain positions, such as dhps-436, dhps-581, and dhps-613, is described as less clear compared to treatment failure, due to their low prevalence in Africa. One can explain why these mutations have received less attention [35]. A study carried out in the Democratic Republic of Congo showed that mutations in dhps-437 and dhps-540 were strongly associated with treatment failure, and this relationship depended on parasitaemia [35]. Patients carrying the dhps (437, 540) mutation had an absolute risk of therapeutic failure on day 14 that was 19% higher than that of subjects without an y mutation, according to this study. In contrast, subjects with high parasitaemia had an 80% higher absolute risk of treatment failure than subjects with the dhps mutation. High parasitaemia could be a sign of low partial immunity, and the greater effect of these mutations associated with high parasitaemia could be due to the immune system’s inability to eliminate resistant parasites. These results therefore suggest that resistance may have a greater influence in less immune patients [35], particularly in children under the age of five. However, due to the evolution of genomic tools, nowadays, those above this age and with the dhfr and dhps mutations should be considered of interest. The WHO recommendation anticipated alternative drugs for intermittent preventive treatment in case the prevalence of Pfdhps 540E mutant parasites exceed 50%. Countries in East and Central Africa are now grappling with a high prevalence of triple mutants. The absence of quintuple mutations, therefore, confirms that SMC still remains an appropriate malaria chemoprevention tool in children under five years of age during the period of high transmission in Burkina Faso. However, high-frequency individual mutations such as Pfdhfr59 threaten the efficacy of the sulfadoxine–pyrimethamine combination, which is also the combination used for intermittent preventive treatment of women in the last two trimesters of pregnancy.
5. Conclusions
This study on parasite diversity and resistant molecular assessment during SMC allowed us to provide data indicating a high genetic diversity of the parasite by the analysis of allelic families with a predominance of the Ro33 allele of msp-1 and 3D7 for msp-2. A reduction in the multiplicity and polyclonality of infection, reflecting a decrease in parasite polymorphism during SMC, was observed, thus indicating a positive effect of this strategy in the treatment and prevention of malaria in children. These data also show a trend towards a reduction in the complexity of infection associated with the level of malaria transmission, which could be considered a beneficial effect of SMC in the children from Saponé comprising the study population. The prevalence of mutations in the genes (Pfcrt and Pfmdr1) associated with parasite resistance to AQ and in the genes (Pfdhfr and Pfdhps) associated with resistance to SP remained low or negligible or non-existent during SMC. This shows that the combination molecules used during SMC (SP+AQ) remain appropriate for the prevention of malaria in children under five during periods of high transmission. Finally, our study once again demonstrates the importance of molecular tools in malaria surveillance, as well as the importance of considering malaria molecular surveillance as a priority to accompany malaria control and elimination.
Author Contributions
I.S., J.S., O.O., and F.W.D. designed and produced the study. I.S., S.N., S.S. (Salif Sombié), S.S.S., F.W.D. and A.A.Z. wrote the original draft. S.N., S.S. (Salif Sombié), S.S.S., N.O (Nicolas Ouedraogo), and S.S. (Salam Sawadogo): carried out the sample collection in the field and the molecular analyses in the laboratory. I.S. and S.N. carried out the data analysis. I.S., J.S., A.B., F.W.D., A.A.Z., and O.O. supervised, revised, and contributed to the manuscript. All authors, including H.S., C.T., R.I., G.Y.N.T., D.O.A.Z., J.N.Y., F.C.A.K., D.Z., A.A.M., and A.Z., have read and agreed to the published version of the manuscript.
Funding
The research was not funded by an external funding company; rather, it was personally funded by the authors.
Informed Consent Statement
Written informed consent was obtained from participants and the parents or legal guardians of all child participants before enrolment. All health concerns were treated free of charge according to the local standards of care.
Acknowledgments
We are grateful to all participants and parents or legal guardians of all the children who took part in this study. We also thank the staff of the Centre National de Recherche et de Formation sur le Paludisme, CNRFP, and the community health agents who contributed to the realization of this study.
Conflicts of Interest
The authors declare that they have no competing interests. There are no conflicts of interest between the authors and producers of the products because we only intend to use these products for the advancement of knowledge.
References
- Cox-Singh, J.; Hiu, J.; Lucas, S.B.; Divis, P.C.; Zulkarnaen, M.; Chandran, P.; Wong, K.T.; Adem, P.; Zaki, S.R.; Singh, B.; et al. Severe malaria—A case of fatal Plasmodium knowlesi infection with post-mortem findings: A case report. Malar. J. 2010, 9, 10. [Google Scholar] [CrossRef]
- WHO. World Malaria World Malaria Report Report. 2023. Available online: https://www.wipo.int/amc/en/mediation/%0Ahttps://www.who.int/teams/global-malaria-programme/reports/world-malaria-report-2023 (accessed on 20 September 2025).
- WHO. Updated WHO Recommendations for Malaria Chemoprevention and Elimination. Global Malaria Programme; WHO: Geneva, Switzerland, 2022. [Google Scholar]
- Oumar, A.A.; Ba, M.; Diallo, D.; Sanogo, M.; Ousmane, F.W.; Diawara, S.I.; Guindo, J.B.; Malan, K.I.; Traore, S.F.; Diop, S. Impact of seasonal malaria chemoprevention in children aged 3 to 59 months in the Kangaba and Kolokani health districts between 2013 and 2015. Rev. Malienne D’infectiol. Microbiol. 2020, 15, 54–60. [Google Scholar] [CrossRef]
- Druetz, T.; Corneau-Tremblay, N.; Millogo, T.; Kouanda, S.; Ly, A.; Bicaba, A.; Haddad, S. Impact Evaluation of Seasonal Malaria Chemoprevention under Routine Program Implementation: A Quasi-ExperimenItal Study in Burkina Faso. Am. J. Trop. Med. Hyg. 2018, 98, 524–533. [Google Scholar] [CrossRef]
- Ministère de la Santé du Burkina Faso. Annuaire Statistique 2018 du Minitère de la Santé; Ministère de la santé du Burkina Faso: Ouagadougou, Burkina Faso, 2019; p. 494. [Google Scholar]
- Somé, A.F.; Bazié, T.; Zongo, I.; Yerbanga, R.S.; Nikiéma, F.; Neya, C.; Taho, L.K.; Ouédraogo, J.-B. Plasmodium falciparum msp1 and msp2 genetic diversity and allele frequencies in parasites isolated from symptomatic malaria patients in Bobo-Dioulasso, Burkina Faso. Parasites Vectors 2018, 11, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Berzosa, P.; Esteban-Cantos, A.; García, L.; González, V.; Navarro, M.; Fernández, T.; Romay-Barja, M.; Herrador, Z.; Rubio, J.M.; Ncogo, P.; et al. Profile of molecular mutations in pfdhfr, pfdhps, pfmdr1, and pfcrt genes of Plasmodium falciparum related to resistance to different anti-malarial drugs in the Bata District (Equatorial Guinea). Malar. J. 2017, 16, 1–10. [Google Scholar] [CrossRef]
- Driss, A.; Hibbert, J.M.; O Wilson, N.; A Iqbal, S.; Adamkiewicz, T.V.; Stiles, J.K. Genetic polymorphisms linked to susceptibility to malaria. Malar. J. 2011, 10, 271. [Google Scholar] [CrossRef] [PubMed]
- Bogreau, H.; Renaud, F.; Bouchiba, H.; Durand, P.; Assi, S.-B.; Henry, M.-C.; Garnotel, E.; Pradines, B.; Fusai, T.; Wade, B.; et al. Genetic diversity and structure of African Plasmodium falciparum populations in urban and rural areas. Am. J. Trop. Med. Hyg. 2006, 74, 953–959. [Google Scholar] [CrossRef]
- Chekol, T.; Alemayehu, G.S.; Tafesse, W.; Legesse, G.; Zerfu, B.; File, T.; Wolde, M.; Golassa, L. Genetic Diversity of Merozoite Surface Protein-1 and -2 Genes in Plasmodium falciparum Isolates among Asymptomatic Population in Boset and Badewacho Districts, Southern Ethiopia. J. Parasitol. Res. 2022, 2022, 1–8. [Google Scholar] [CrossRef]
- Atroosh, W.M.; Al-Mekhlafi, H.M.; Mahdy, M.A.; Saif-Ali, R.; Al-Mekhlafi, A.M.; Surin, J. Genetic diversity of Plasmodium falciparum isolates from Pahang, Malaysia based on MSP-1 and MSP-2 genes. Parasites Vectors 2011, 4, 233. [Google Scholar] [CrossRef]
- Badoum, E.S.; Bougouma, E.C.; Sombie, S.; Sermé, S.S.; Yaro, J.B.; Diarra, A.; Nébié, I.; Ouedraogo, A.; Tiono, A.B.; Soulama, I.; et al. Relationship between human genetic factors and Plasmodium falciparum genetic diversity of msp1, msp2 and glurp in a malaria endemic area of Burkina Faso. Biomed. Genet. Genom. 2019, 4, 144. [Google Scholar] [CrossRef]
- Soulama, I.; Nébié, I.; Ouédraogo, A.; Gansane, A.; Diarra, A.; Tiono, A.B.; Bougouma, E.C.; Konaté, A.T.; Kabré, G.B.; Taylor, W.R.; et al. Plasmodium falciparum genotypes diversity in symptomatic malaria of children living in an urban and a rural setting in burkina Faso. Malar. J. 2009, 8, 135. [Google Scholar] [CrossRef]
- Mohammed, H.; Kassa, M.; Mekete, K.; Assefa, A.; Taye, G.; Commons, R.J. Genetic diversity of the msp-1, msp-2, and glurp genes of Plasmodium falciparum isolates in Northwest Ethiopia. Malar. J. 2018, 17, 1–8. [Google Scholar] [CrossRef]
- Takala, S.; Escalante, A.; Branch, O.; Kariuki, S.; Biswas, S.; Chaiyaroj, S.; Lal, A. Genetic diversity in the Block 2 region of the merozoite surface protein 1 (MSP-1) of Plasmodium falciparum: Additional complexity and selection and convergence in fragment size polymorphism. Infect. Genet. Evol. 2006, 6, 417–424. [Google Scholar] [CrossRef]
- Sondo, P.; Derra, K.; Lefevre, T.; Diallo-Nakanabo, S.; Tarnagda, Z.; Zampa, O.; Kazienga, A.; Valea, I.; Sorgho, H.; Ouedraogo, J.-B.; et al. Genetically diverse Plasmodium falciparum infections, within-host competition and symptomatic malaria in humans. Sci. Rep. 2019, 9, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Amodu, O.; Olumese, P.; Gbadegesin, R.; Ayoola, O.; Adeyemo, A. The influence of individual preventive measures on the clinical severity of malaria among Nigerian children. Acta Trop. 2006, 97, 370–372. [Google Scholar] [CrossRef]
- Bouyou-Akotet, M.K.; Offouga, C.L.; Mawili-Mboumba, D.P.; Essola, L.; Madoungou, B.; Kombila, M. Falciparum malaria as an emerging cause of fever in adults living in Gabon, Central Africa. BioMed. Res. Int. 2014, 2014, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Ariey, F.; Chalvet, W.; Hommel, D.; Peneau, C.; Hulin, A.; Mercereau-Puijalon, O.; Duchemin, J.B.; Sarthou, J.L.; Reynes, J.M.; Fandeur, T. Plasmodium falciparum parasites in French Guiana: Limited genetic diversity and high selfing rate. Am. J. Trop. Med. Hyg. 1999, 61, 978–985. [Google Scholar] [CrossRef] [PubMed]
- Al-Yaman, F.; Genton, B.; Reeder, J.; Anders, R.; Smith, T.; Alpers, M. Reduced risk of clinical malaria in children infected with multiple clones of Plasmodium falciparum in a highly endemic area: A prospective community study. Trans. R. Soc. Trop. Med. Hyg. 1997, 91, 602–605. [Google Scholar] [CrossRef]
- Sondo, P.; Derra, K.; Nakanabo, S.D.; Tarnagda, Z.; Kazienga, A.; Zampa, O.; Valéa, I.; Sorgho, H.; Owusu-Dabo, E.; Ouédraogo, J.-B.; et al. Artesunate-amodiaquine and artemether-lumefantrine therapies and selection of pfcrt and PfmdR1 alleles in Nanoro, Burkina Faso. PLoS ONE 2016, 11, e0151565. [Google Scholar] [CrossRef]
- Laishram, D.D.; Sutton, P.L.; Nanda, N.; Sharma, V.L.; Sobti, R.C.; Carlton, J.M.; Joshi, H. The complexities of malaria disease manifestations with a focus on asymptomatic malaria. Malar. J. 2012, 11, 29. [Google Scholar] [CrossRef]
- Sodeman, W.A.; Jeffery, G.M. Primary malarial thrombocytopenia in the rhesus monkey. Trans. R. Soc. Trop. Med. Hyg. 1966, 60, 70–74. [Google Scholar] [CrossRef]
- Mockenhaupt, F.P.; Bedu-Addo, G.; von Gaertner, C.; Boyé, R.; Fricke, K.; Hannibal, I.; Karakaya, F.; Schaller, M.; Ulmen, U.; A Acquah, P.; et al. Detection and clinical manifestation of placental malaria in southern Ghana. Malar. J. 2006, 5, 119. [Google Scholar] [CrossRef] [PubMed]
- Sondo, P.; Bihoun, B.; Kabore, B.; Tahita, M.C.; Derra, K.; Rouamba, T.; Diallo, S.N.; Kazienga, A.; Ilboudo, H.; Valea, I.; et al. Polymorphisme de Plasmodium falciparum et mutations des gènes de résistance Pfcrt et Pfmdr1 dans la zone de Nanoro, Burkina Faso. Pan Afr. Med. J. 2021, 39, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Djimdé, A.; Doumbo, O.K.; Cortese, J.F.; Kayentao, K.; Doumbo, S.; Diourté, Y.; Coulibaly, D.; Dicko, A.; Su, X.-Z.; Nomura, T.; et al. A molecular marker for chloroquine-resistant falciparum malaria. N. Engl. J. Med. 2001, 344, 257–263. [Google Scholar] [CrossRef]
- Atroosh, W.M.; Al-Mekhlafi, H.M.; Mahdy, M.A.; Surin, J. The detection of pfcrt and pfmdr1 point mutations as molecular markers of chloroquine drug resistance, Pahang, Malaysia. Malar. J. 2012, 11, 251. [Google Scholar] [CrossRef]
- Salissou, I.; Moustapha, L.M.; Yerima, B.; Alkassoum, I.; Hadiza, D.; Ibrahim, L. Perception de la chimioprévention du paludisme saisonnier au Niger Perception of the seasonal malaria chemoprevention in Niger. Int. J. Biol. Chem. Sci. 2016, 10, 2710–2715. [Google Scholar] [CrossRef]
- Venkatesan, M.; Gadalla, N.B.; Stepniewska, K.; Dahal, P.; Nsanzabana, C.; Moriera, C.; Price, R.N.; Mårtensson, A.; Rosenthal, P.J.; Dorsey, G.; et al. Polymorphisms in Plasmodium falciparum chloroquine resistance transporter and multidrug resistance 1 genes: Parasite risk factors that affect treatment outcomes for P. falciparum malaria after artemether-lumefantrine and artesunate-amodiaquine. Am. J. Trop. Med. Hyg. 2014, 91, 833–843. [Google Scholar] [CrossRef]
- Mahamar, A.; Sumner, K.M.; Levitt, B.; Freedman, B.; Traore, A.; Barry, A.; Issiaka, D.; Dembele, A.B.; Kanoute, M.B.; Attaher, O.; et al. Effect of three years’ seasonal malaria chemoprevention on molecular markers of resistance of Plasmodium falciparum to sulfadoxine-pyrimethamine and amodiaquine in Ouelessebougou, Mali. Malar. J. 2022, 21, 39. [Google Scholar] [CrossRef]
- Baba, E.; Hamade, P.; Kivumbi, H.; Marasciulo, M.; Maxwell, K.; Moroso, D.; Roca-Feltrer, A.; Sanogo, A.; Johansson, J.S.; Tibenderana, J.; et al. Effectiveness of seasonal malaria chemoprevention at scale in west and central Africa: An observational study. Lancet 2020, 396, 1829–1840. [Google Scholar] [CrossRef]
- Cairns, M.E.; Sagara, I.; Zongo, I.; Kuepfer, I.; Thera, I.; Nikiema, F.; Diarra, M.; Yerbanga, S.R.; Barry, A.; Tapily, A.; et al. Evaluation of seasonal malaria chemoprevention in two areas of intense seasonal malaria transmission: Secondary analysis of a household-randomised, placebo-controlled trial in Houndé District, Burkina Faso and Bougouni District, Mali. PLoS Med. 2020, 17, e1003214. [Google Scholar] [CrossRef]
- Rosillo, S.R.; Dimbu, P.R.; Cândido, A.L.M.; Oh, J.H.M.; Ferreira, C.M.; Andrade, B.N.; Labuda, S.; Horth, R.; Kelley, J.; Morais, J.F.M.; et al. Molecular Markers of Sulfadoxine-Pyrimethamine Resistance in Samples from Children with Uncomplicated Plasmodium falciparum at Three Sites in Angola in 2019. Antimicrob. Agents Chemother. 2023, 67, e0160122. [Google Scholar] [CrossRef]
- Alker, A.P.; Kazadi, W.M.; Kutelemeni, A.K.; Bloland, P.B.; Tshefu, A.K.; Meshnick, S.R. dhfr and dhps genotype and sulfadoxine-pyrimethamine treatment failure in children with falciparum malaria in the Democratic Republic of Congo. Trop. Med. Int. Health 2008, 13, 1384–1391. [Google Scholar] [CrossRef] [PubMed]
© 2026 Copyright by Authors. Licensed as an open access article using a CC BY 4.0 license.


